Welcome to the neuroLIT Documentation!
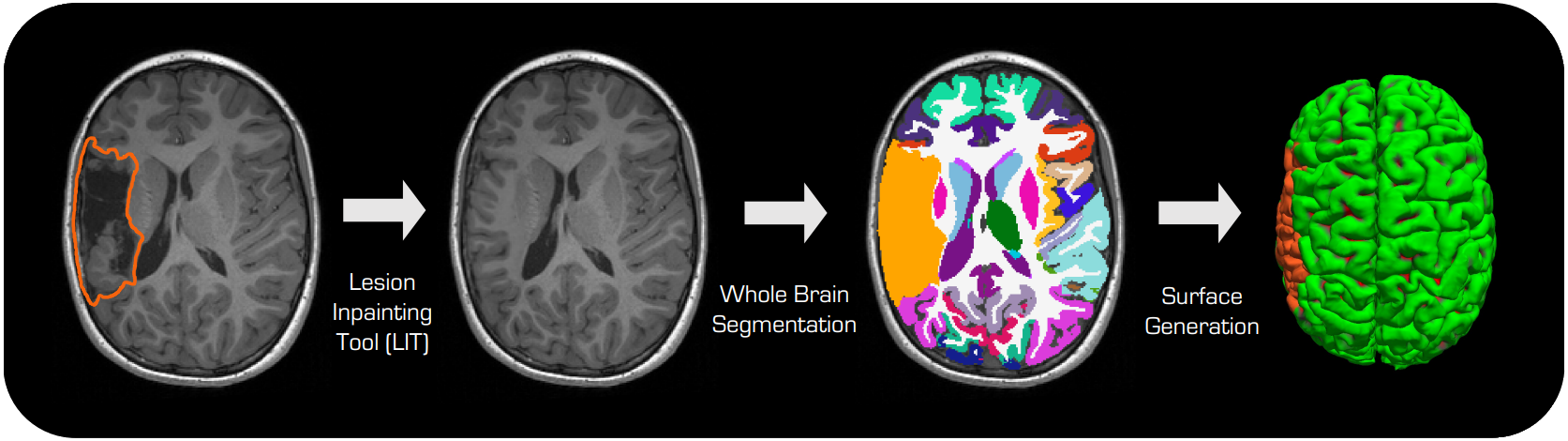
neuroLIT (Neuro Lesion Inpainting Tool) is a tool for inpainting lesions in brain MRI images, independent of their shape or appearance, for further downstream analysis.
neuroLIT with FastSurfer can be run by directly running FastSurfer. For other tools and FreeSurfer use the neuroLIT repository.
🔥 Key Features
Inpaints lesions of any shape or appearance in T1-weighted MRI images
Standalone operation or integration with FastSurfer
Docker and Singularity containerization support
PyPI package available for easy installation
Surface and segmentation masking capabilities
Quick Start
Using Docker/Singularity (Recommended)
git clone https://github.com/Deep-MI/neurolit.git && cd neurolit
./neurolit/scripts/run_lit_containerized.sh --input_image T1w.nii.gz \\
--mask_image lesion_mask.nii.gz --output_directory output_directory
Using PyPI Package
# Install the package
pip install neurolit
# Download model checkpoints
lit-download-models
# Run LIT
lit-inpainting --input_image T1w.nii.gz --lesion_mask lesion_mask.nii.gz \\
--output_directory output_directory --dilate 2
User Guide
- Installation
- Usage Guide
- Downstream Analysis with Lesion Exclusion
- Surface-Based Analysis
- Step 1 — Obtain Lesion Surface Labels
- Step 2 — Extract Regional Statistics (Optional)
- Step 3 — Project to Common Surface Template
- Step 4 — Mask Lesion Vertices
- Step 5 — Run Your Statistical Analysis
- Step 6 — Visualize on the Cortical Surface (Optional)
- Volumetric Analysis
- Dependencies
- Training
Development
Reference